Table of Contents
Definition / general | Essential features | Terminology | ICD coding | Epidemiology | Sites | Pathophysiology | Etiology | Diagrams / tables | Diagnosis | Laboratory | Prognostic factors | Case reports | Treatment | Microscopic (histologic) description | Microscopic (histologic) images | Virtual slides | Peripheral smear description | Peripheral smear images | Positive stains | Negative stains | Flow cytometry description | Flow cytometry images | Molecular / cytogenetics description | Molecular / cytogenetics images | Sample pathology report | Differential diagnosis | Additional references | Board review style question #1 | Board review style answer #1 | Board review style question #2 | Board review style answer #2Cite this page: Morrow WP, Insuasti-Beltran G. AML with other KMT2A rearrangements. PathologyOutlines.com website. https://www.pathologyoutlines.com/topic/bonemarrowneoplasticAMLKMT2A.html. Accessed April 2nd, 2025.
Definition / general
- Acute myeloid leukemia (AML) with other KMT2A rearrangements: subclassification of AML defined by the International Consensus Classification (ICC) as AML with rearrangement of the KMT2A gene other than t(9;11) / MLLT3::KMT2A
- Recurrent translocations recognized in this category include AMLs with t(4;11) AFF1::KMT2A; t(6;11) / AFDN::KMT2A; t(10;11) / MLLT10::KMT2A; t(10;11) / TET1::KMT2A; t(11;19) / KMT2A::ELL; t(11;19) / KMT2A::MLLT1
- In the 5th edition of WHO, the category of AML with KMT2A rearrangement includes any KMT2A gene rearrangement, regardless of the partner gene
Essential features
- KMT2A rearrangement is a recurrent genetic finding identified in both acute lymphoblastic leukemias / lymphomas (ALL) and AML
- KMT2A rearrangement has an inverse association with patient age, with a large subset of infantile cases of AML presenting with KMT2A rearrangement
- AML with KMT2A rearrangement most commonly presents with a monocytic phenotype, with myelomonocytic and myeloid phenotypes less frequent
- KMT2A has numerous partner genes; by International Consensus Classification (ICC) 2022 definition, AML with other KMT2A rearrangements include t(4;11) AFF1::KMT2A; t(6;11) / AFDN::KMT2A; t(10;11) / MLLT10::KMT2A; t(10;11) / TET1::KMT2A; t(11;19) / KMT2A::ELL; t(11;19) / KMT2A::MLLT1
- Overall prognosis is poor but prognosis varies somewhat with fusion gene
- KMT2A rearrangement can be identified by various testing methods including conventional karyotype, FISH, PCR and NGS
Terminology
- KMT2A gene was previously named mixed lineage leukemia (MLL)
- WHO 5th edition classification: acute myeloid leukemia with KMT2A rearrangement
- ICC 2022 classification: acute myeloid leukemia with other KMT2A rearrangements
ICD coding
- ICD-O: 9897/3 - acute myeloid leukemia with KMT2A rearrangement
- ICD-10
- ICD-11: 2A60.0 & XH1E41 - acute myeloid leukemia with recurrent genetic abnormalities & acute myeloid leukemia, 11q23 abnormalities
Epidemiology
- KMT2A rearrangement
- Incidence has an inverse relationship with age (Blood 2003;102:2395)
- Most frequently encountered genetic abnormality in pediatric population, accounting for 20% of pediatric AML cases
- Less common among adult population, accounting for 2 - 3% of adult AML cases
- Has increased prevalence among patients with AML post-cytotoxic therapy (AML pCT) (Cytometry B Clin Cytom 2022;102:123)
Sites
- By definition, AML with other KMT2A rearrangement involves bone marrow and peripheral blood
- Prevalence of extramedullary disease is higher than that of non-KMT2A rearranged leukemia in both adults and pediatric populations with a high frequency of fusion partners (Proc Natl Acad Sci U S A 2020;117:26340)
- Extramedullary disease most frequently manifests in the liver, spleen or CNS; less commonly as myeloid sarcoma in other locations
- Data is limited for low frequency fusion partners
Pathophysiology
- Myeloid precursor cell undergoes genetic event resulting in rearrangement of KMT2A gene on chromosome 11
- KMT2A rearrangement in AML usually manifests as a translocation
- Rearrangement of KMT2A disrupts its usual role in regulating gene expression during early development and hematopoiesis; this disruption will lead to clonal expansion of the myeloid cell lineage harboring the KMT2A (Cancer Lett 2019;458:56)
- Clonal expansion will result in increased myeloid lineage blasts, further developing into bone marrow replacement and subsequent pancytopenias
Etiology
- Definitive etiology is not well understood
- Risk factors in adult patients include cytotoxic chemotherapy and other nonspecific DNA damaging events
Diagnosis
- WHO criteria for AML with KMT2A rearrangement
- Essential
- Myeloid neoplasm with increased peripheral blood or bone marrow blasts (may be < 20%)
- Blasts express a myeloid immunophenotype, not fulfilling immunophenotypic criteria for mixed phenotype AML
- Presence of KMT2A rearrangement
- Not fulfilling diagnostic criteria for myeloid neoplasm post-cytotoxic therapy
- Desirable
- Identification of the KMT2A fusion partner
- ICC 2022 criteria for AML with other KMT2A rearrangement (Blood 2022;140:1200)
- Overall criteria similar to the WHO, except importantly t(9;11) / MLLT3::KMT2A is categorized as a separate entity
- Recognized recurrent fusion partners are described in other sections
- Identification of KMT2A rearrangement is critical for this diagnosis (see Molecular / cytogenetics description for further description)
- At least 10% myeloblasts in the peripheral blood or bone marrow sampling are required for diagnosis by ICC classification
- Essential
Laboratory
- Anemia and thrombocytopenia likely present at diagnosis
- WBC will vary case to case; ranges from leukopenia to leukocytosis and correlates with number of circulating blasts
Prognostic factors
- In general, prognosis of KMT2A rearranged AML is poor with high risk of relapse; however, prognosis varies upon fusion partner to KMT2A gene (Blood Cancer J 2021;11:162)
- Few partner genes may give intermediate to good prognosis
- Overall median survival in patients with KMT2A rearrangement is 0.9 years
- Additional cytogenetic abnormalities and mutations, especially in myelodysplasia associated genes, worsens prognosis
Case reports
- 31 weeks gestational age female neonate with AFF1::KMT2A fusion AML (Oncol Lett 2022;24:283)
- 6 year old girl with KMT2A::MLLT10 fusion AML (Cancer Genet 2023;276:36)
- 7 year old boy with KMT2A::MLLT1 fusion AML (Cytogenet Genome Res 2019;157:213)
- 43 year old woman with KMT2A::SEPT5 fusion AML (SAGE Open Med Case Rep 2018;6:2050313X17750334)
- 63 year old woman with hepatic gastrointestinal stromal tumor (GIST) and concomitant KMT2A::ELL fusion AML (World J Clin Cases 2020;8:1251)
Treatment
- Intensive chemotherapy as induction followed by additional consolidation therapy
- Trials utilizing novel agents such as MEK inhibitors, demethylating agents and other targeted therapies are ongoing (Front Pharmacol 2022;13:749472)
- Hematopoietic stem cell transplant
Microscopic (histologic) description
- Increased myeloid lineage blasts (percentage will vary case to case)
- WHO 5th edition does not have a blast requirement and the ICC requires at least 10% blasts in peripheral blood or bone marrow
- On the aspirate smears, crush prep or touch prep, blast morphology will vary by phenotype
- Myeloblasts will have open chromatin, high N:C ratio, prominent nucleoli and possible Auer rods in the cytoplasm
- Monoblasts will have open chromatin, lower N:C ratio, possible cytoplasmic vacuoles and larger cell size
- AML with monocytic differentiation tends to have a spectrum of monocyte differentiation, with mature monocytes, promonocytes and monoblasts (Haematologica 2009;94:994)
- Unless there are coinciding myelodysplasia related gene mutations, significant dysplasia is not expected
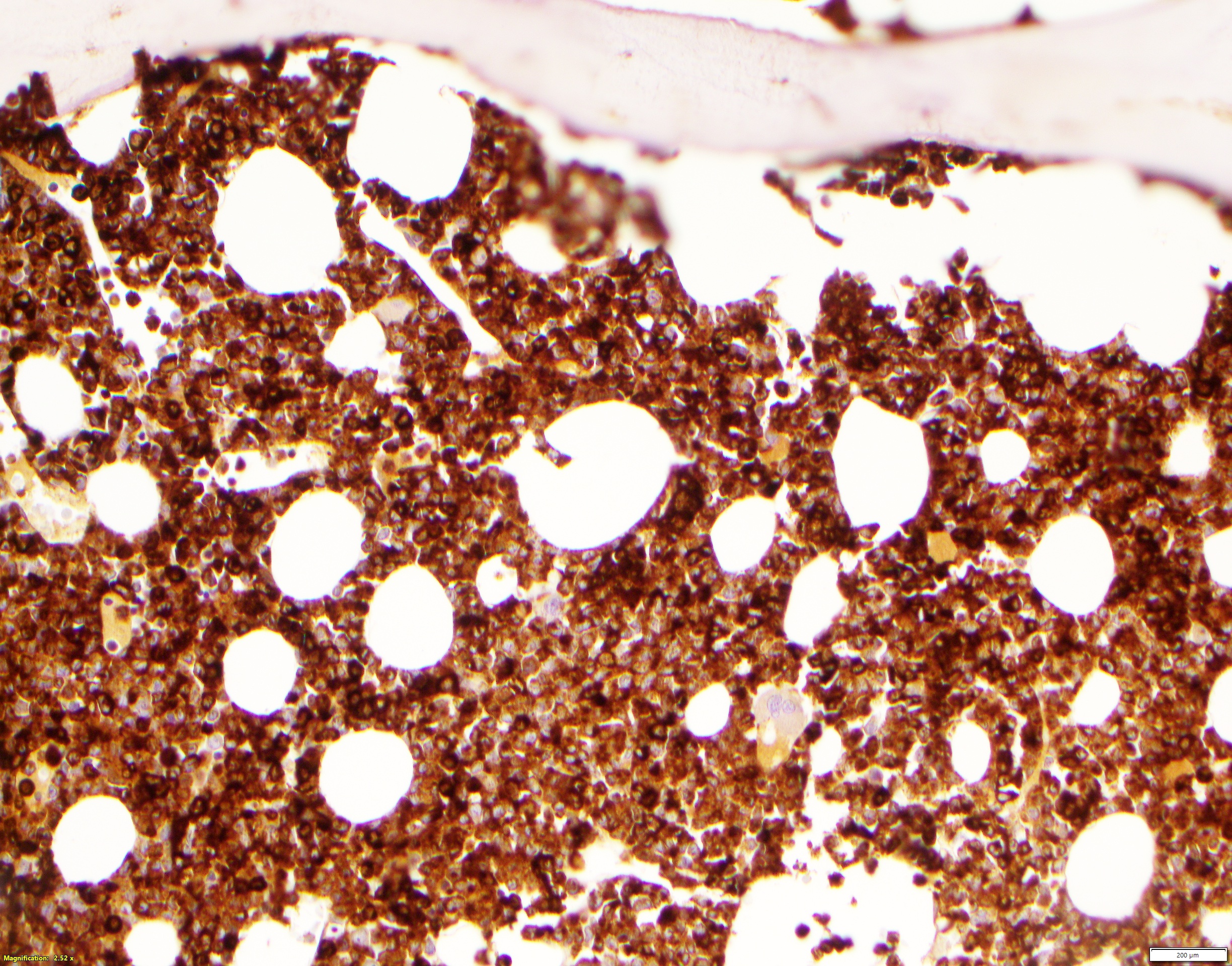
- On core biopsies, varying degrees of bone marrow involvement by blasts will be seen
- Blasts will appear as immature cells with intermediate to large cell size, open chromatin and usually with prominent nucleoli
- Pattern of involvement may be interstitial, aggregate or sheets
- Degree of bone marrow effacement by leukemia will impact background trilineage hematopoiesis
- Trilineage hematopoiesis will be affected by treatment in follow up bone marrow biopsies
Microscopic (histologic) images
Peripheral smear description
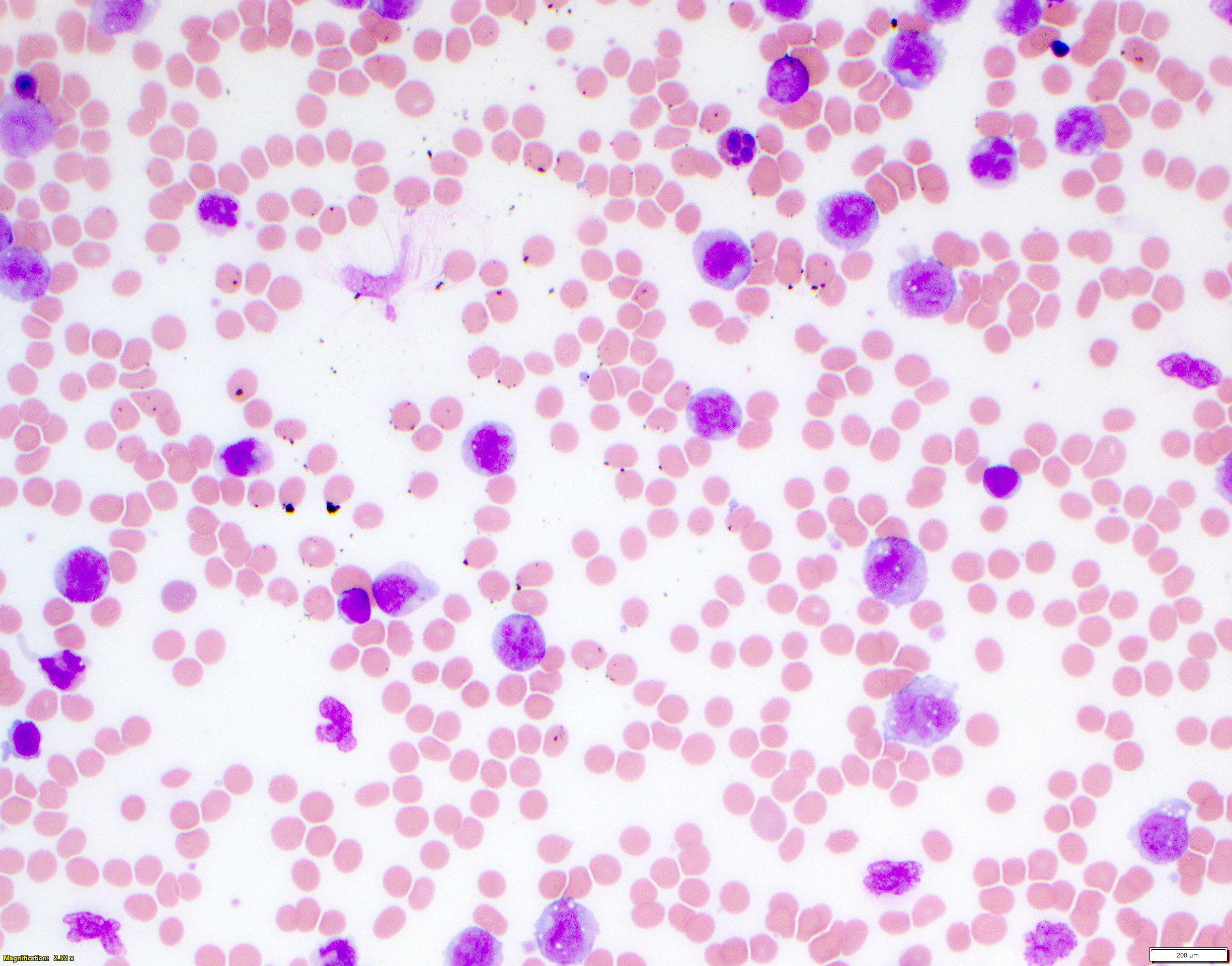
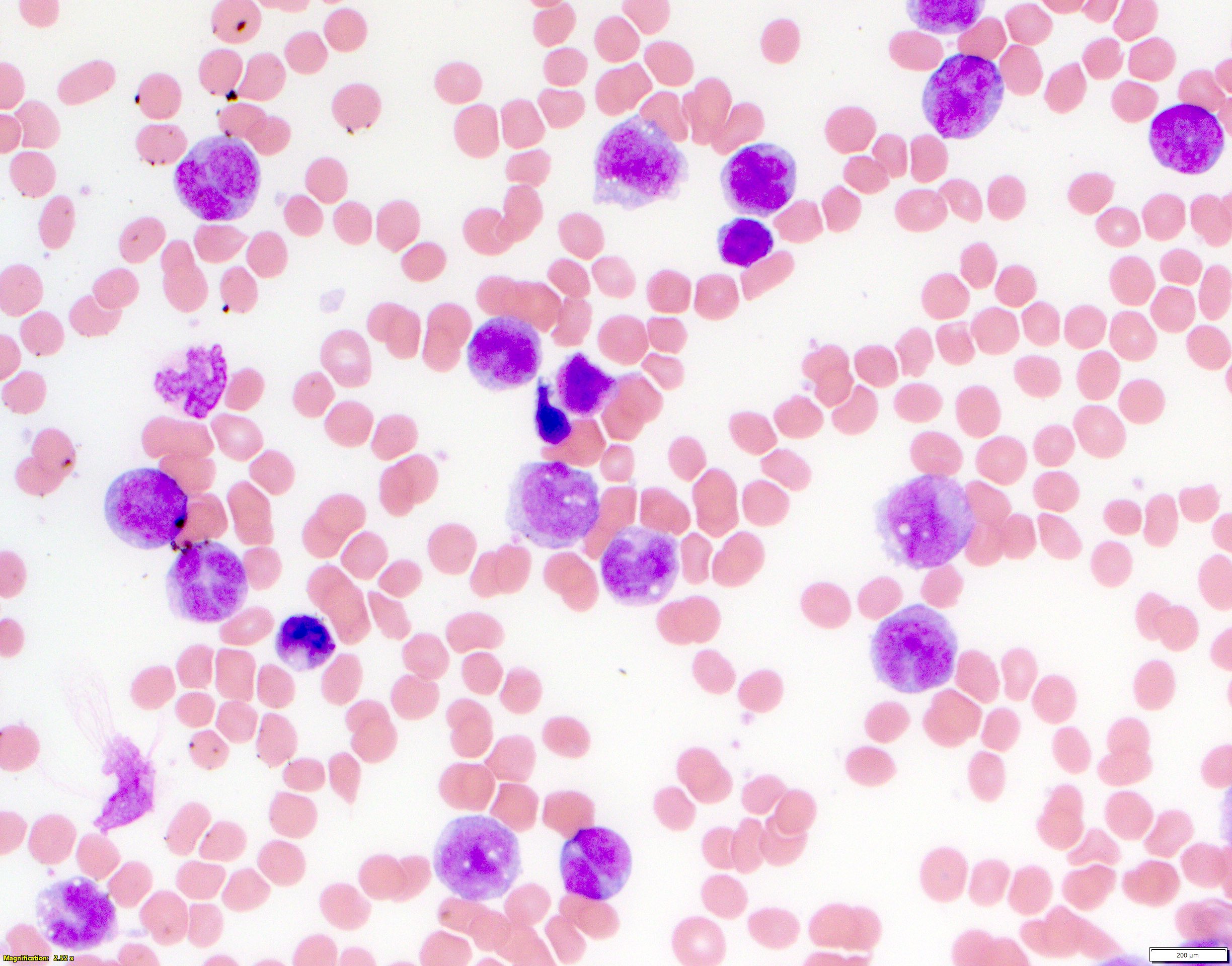
- Increased circulating blasts, frequently with monocytic features, such as increased cytoplasm, vacuoles, cleaved or grooved nuclei
- Variable degree of cytopenias depending on level of bone marrow involvement
- Typically a normocytic, normochromic anemia with otherwise unremarkable red blood cell morphology
- Typically a thrombocytopenia with unremarkable platelet morphology
- No features of dysplasia in granulocytes unless there is a coinciding myelodysplasia related gene mutation(s)
- Reference: Haematologica 2009;94:994
Peripheral smear images
Positive stains
Negative stains
Flow cytometry description
- Immunophenotype by flow cytometry varies but most commonly represents monocytic differentiation, with a smaller subset of cases having myelomonocytic or myeloid immunophenotype
- Cases with monocytic differentiation will be positive for CD33, CD13, CD64, CD4, HLA-DR, CD14 (variable) and negative for CD34, CD117 and MPO
- Cases with myeloid differentiation will be positive for CD34, MPO, CD33, HLA-DR and negative for CD64, CD4 and CD14
- CD123 is positive in 90% of cases with KMT2A rearrangement (Cytometry B Clin Cytom 2022;102:123)
- Markers of lymphoblastic differentiation, such as TdT, CD3, CD19, CD10 are almost universally negative
Flow cytometry images
Molecular / cytogenetics description
- KMT2A gene (lysine methyltransferase 2A) is located on chromosome 11q23.3
- More than 130 different partners have been recognized in association with KMT2A; AFF1, MLLT1, MLLT3, MLLT10, MLLT4 and ELL are the most frequently recognized partners in order of decreasing frequency (BMC Med Genomics 2020;13:106)
- These genes represent over 90% of KMT2A rearrangement cases and are specifically recognized by the ICC 2022 classification for KMT2A with other rearrangement
- Due to the various different breakpoints in the KMT2A gene, comprehensive testing for KMT2A rearrangement can be challenging
- Different types of structural rearrangements can be the underlying cause of KMT2A fusions, including insertions, deletions, inversions and translocations
- Several different molecular and cytogenetic testing modalities can identify KMT2A rearrangement
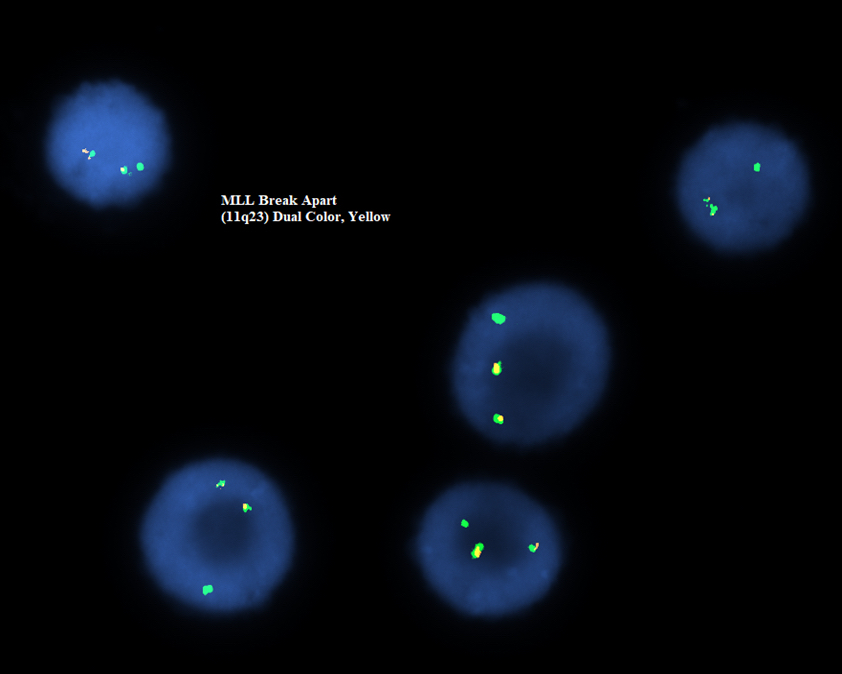
- FISH is routinely performed using the aspirate material collected during the bone marrow biopsy procedure; KMT2A break apart probes (with a probe labeling 3’KMT2A and a probe labeling 5’KMT2A) are routinely ordered as part of a FISH panel for a diagnostic leukemia marrow
- Some rearrangements may be cryptic by FISH, such as KMT2A::USP2
- Routine karyotype can detect many translocations involved in KMT2A rearrangement (such as t(9;11)); however, some translocations may be cryptic (such as t(10;11) MLLT10::KMT2A)
- Additional cytogenetic alterations can coincide with KMT2A rearrangement
- PCR can be used to detect specific fusion transcripts
- Some available panels include up to 20 fusion genes, including all the recurrent fusions identified
- Myeloid next generation sequencing (NGS) panels may have KMT2A gene incorporated into a fusion panel and can detect KMT2A
- FISH is routinely performed using the aspirate material collected during the bone marrow biopsy procedure; KMT2A break apart probes (with a probe labeling 3’KMT2A and a probe labeling 5’KMT2A) are routinely ordered as part of a FISH panel for a diagnostic leukemia marrow
- Of note, partial tandem duplication (PTD) of KMT2A may be detected by NGS; PTD of the KMT2A gene does not qualify as KMT2A rearrangement for either classification scheme
Molecular / cytogenetics images
Sample pathology report
- Bone marrow, biopsy:
- Acute myeloid leukemia with other KMT2A rearrangement (see comment)
- Comment: Increased blasts (50%) in a hypercellular marrow (90%). The blasts are morphologically and immunophenotypically consistent with myeloblasts. Concurrent FISH studies identified KMT2A rearrangement (see FISH report for full details). Concurrent karyotype studies identified t(10;11) (see karyotype report for full details). A myeloid NGS panel recognized a KMT2A::MLLT10. Overall, these findings are consistent with acute myeloid leukemia with KMT2A rearrangement (WHO 5th edition) or acute myeloid leukemia with other KMT2A rearrangement (ICC 2022).
Differential diagnosis
- Acute myeloid leukemia with t(9;11) / MLLT3::KMT2A:
- Differentiated by defining KMT2A rearrangement
- Classified separately by the ICC
- Shares classification by the WHO 5th edition
- Acute myeloid leukemia with other defining genetic abnormalities:
- Lacks KMT2A rearrangement
- Presence of other defining genetic feature
- Acute lymphoblastic leukemia with KMT2A rearrangement:
- Differentiated by immunophenotype and cell lineage
- Acute monocytic leukemia:
- Myeloid leukemia with monocytic features lacking KMT2A rearrangement
- Chronic myelomonocytic leukemia with KMT2A rearrangement:
- No increase in blasts
- Dysplastic features
- Rare and difficult to diagnose
- Increased blasts and KMT2A rearrangement better classified as AML
Additional references
Board review style question #1
A patient with newly diagnosed acute myeloid leukemia had cytogenetics, FISH and myeloid NGS testing performed. FISH identified findings seen in the above image and myeloid NGS identified a KMT2A::AFDN fusion. Per the ICC 2022 guidelines, which of the following is the best classification of this patient's disease?
- Acute myeloid leukemia, NOS
- Acute myeloid leukemia with myelodysplasia related cytogenetics
- Acute myeloid leukemia with other KMT2A translocation
- Acute myeloid leukemia with t(9;11) / MLLT3::KMT2A
- B lymphoblastic leukemia with t(v;11q23.3) / KMT2A rearrangement
Board review style answer #1
C. Acute myeloid leukemia with other KMT2A translocation. The t(6;11) translocation (KMT2A::AFDN) is recognized by the ICC 2022 classification as one of the recurring KMT2A rearrangements other than MLLT3::KMT2A. Answer A is incorrect because this acute myeloid leukemia has a defining genetic alteration. Answer B is incorrect because the t(6;11) translocation (KMT2A::AFDN) is not recognized as a myelodysplasia related cytogenetic change or gene mutation. Answer D is incorrect because that is not the identified molecular alteration in this case. Answer E is incorrect because the question stem described this case as a myeloid leukemia, as well as a different molecular alteration.
Comment Here
Reference: AML with other KMT2A rearrangements
Comment Here
Reference: AML with other KMT2A rearrangements
Board review style question #2
Acute myeloid leukemia with KMT2A rearrangement most commonly presents with what differentiation?
- Eosinophilic
- Lymphoblastic
- Monocytic
- Myeloid
- Myelomonocytic
Board review style answer #2
C. Monocytic. Acute myeloid leukemias with KMT2A rearrangement most commonly present with a monocytic immunophenotype. Answer A is incorrect because acute myeloid leukemia with KMT2A rearrangement does not present with eosinophilic differentiation. Answer B is incorrect because while lymphoblastic leukemias can present with KMT2A rearrangements, the question was asking for differentiation of myeloid leukemias. Answers D and E are incorrect because although acute myeloid leukemia with KMT2A rearrangement can present with myeloid or myelomonocytic differentiation, it is far more common to have pure monocytic differentiation.
Comment Here
Reference: AML with other KMT2A rearrangements
Comment Here
Reference: AML with other KMT2A rearrangements